5.1. One- and two-dimensional PDFs
[Note: If there are questions in this section you can’t answer right away, come back to them after playing with 📥 Demo: Exploring PDFs.]
To a Bayesian, everything is a PDF
Physicists are used to dealing with PDFs as normalized wave functions squared. For a one-dimensional particle, the probability density at \(x\) is
\[
|\Psi(x)|^2 \Longrightarrow p(x)
\]
The probability of finding \(x\) in some interval \(a \leq x \leq b\) is found by integration:
\[
\mathbb{P}(a \leq x \leq b) = \int_a^b |\Psi(x)|^2 \; dx
\]
Just as with “Lagrangian” vs. “Lagrangian density”, physicists are not always careful when saying probability vs. probability density.
Physicists are also used to multidimensional normalized PDFs as wave functions squared, e.g. probability density for particle 1 at \(x_1\) and particle 2 at \(x_2\):
\[
|\Psi(x_1, x_2)|^2 \Longrightarrow p(x_1,x_2) \equiv p(\textbf{x})
\quad \mbox{with}\quad \textbf{x}
\equiv \{x_1,x_2\}
\]
(Note that you will find alternative notation in the physics and statistics literature for generic PDFs: \(p(\textbf{x}) = P(\textbf{x}) = \textrm{pr}(\textbf{x}) = \textrm{prob}(\textbf{x}) = \ldots\) )
Some other vocabulary and definitions to remember
\(p(x_1,x_2)\) is the joint probability density of \(x_1\) and \(x_2\).
The marginal probability density of \(x_1\) is:
\(p(x_1) = \int\! p(x_1,x_2)\,dx_2\).
In quantum mechanics, this corresponds to the probability to find particle 1 at \(x_1\) and particle 2 anywhere; that is, \(\int\! |\Psi(x_1,x_2)|^2 dx_2\) (integrated over the full domain of \(x_2\), e.g., 0 to \(\infty\)).
“Marginalizing” = “integrating out” (eliminates from the posterior the “nuisance parameters” whose value you don’t care about, at least not at that moment).
In Bayesian statistics there are PDFs (or PMFs if discrete) for experimental and theoretical uncertainties, model parameters, hyperparameters (what are those?), events (“Will it rain tomorrow?”), etc. Even if \(x\) has the definite value \(x_0\), we can represent this knowledge with a PDF \(p(x) = \delta(x-x_0)\).
Characteristics of PDFs
What are the characteristics (e.g., symmetry, heavy tails, …) of different PDFs: normal, beta, student t, \(\chi^2\), \(\ldots\)
Answer
Check these yourself! Note that the answers will often depend on the parameter values for the distribution. A Student t distribution may have “heavy tails” (meaning more probability in the tails than a Gaussian would have) for some parameter values but for others it approaches a normal distribution (so by definition no heavy tails).
Sampling
What does sampling mean?
Answer
To sample a given distribution is to draw values that represent it. The probability to draw a specific value is given by the distribution. In Bayesian inference one is most often sampling a posterior distribution.
Verifying sampling
How do you know if a distribution is correctly sampled?
Answer
One way is to look at the (normalized) histogram. If correctly sampled, this should approximate the distribution and the approximation should improve with more samples.
Visualizing correlated Gaussian distributions
A multivariate Gaussian distribution for \(N\) dimensional \(\boldsymbol{x} = \{x_1, \ldots, x_N\}\) with \(\boldsymbol{\mu} = \{\mu_1, \ldots, \mu_N\}\), with positive-definite covariance matrix \(\Sigma\) is
\[
p(\boldsymbol{x}|\boldsymbol{\mu},\Sigma) = \frac{1}{\sqrt{\det(2\pi\Sigma)}}
e^{-\frac{1}{2}(\boldsymbol{x}-\boldsymbol{\mu})^\intercal\Sigma^{-1}(\boldsymbol{x}-\boldsymbol{\mu})}
\]
For the one dimensional case, it reduces to the familiar
\[
p(x_1|\mu_1,\sigma_1) = \frac{1}{\sqrt{2\pi\sigma_1^2}}
e^{-\frac{(x_1-\mu_1)^2}{2\sigma_1^2}}
\]
with \(\Sigma = \sigma_1^2\).
For the bivariate case (two dimensional),
\[\begin{split}
\boldsymbol{x} = \pmatrix{x_1\\ x_2} \quad\mbox{and}\quad
\boldsymbol{\mu} = \pmatrix{\mu_1\\ \mu_2} \quad\mbox{and}\quad
\Sigma = \pmatrix{\sigma_1^2 & \rho_{12} \sigma_1\sigma_2 \\
\rho_{12}\sigma_1\sigma_2 & \sigma_2^2}
\quad\mbox{with}\ 0 < \rho_{12}^2 < 1
\end{split}\]
and \(\Sigma\) is positive definite.
Widget user interface features:
Set the mean position \((\mu_1, \mu_2)\) and variances \((\Sigma_{11}, \Sigma_{22})\) with the sliders
Set the correlation \(\rho_{12}\) with the slider. This controls the covariance \(\Sigma_{12} = \rho_{12} \sqrt{ \Sigma_{11} \Sigma_{22}}\).
Four presets are available.
The corner plot shows samples from the bivariate PDF and histograms for the two marginal distributions. Control the number of samples with the slider.
Dashed lines on the marginals mark the 16th, 50th, and 84th percentiles. This is equivalent to \(\pm 1\sigma\) for a one-dimensional Gaussian.
The solid and dashed ellipses in the joint panel are iso-probability levels. These correspond to fixed values of the squared Mahalanobis distance
\[
\Delta^2 = (\mathbf{x} - \boldsymbol{\mu})^\top \Sigma^{-1} (\mathbf{x} - \boldsymbol{\mu}).
\]
In the widget they are drawn at \(\Delta = 1\) and \(\Delta = 2\). Their semi-axes are aligned with the eigenvectors of \(\Sigma\) and have lengths \(\sqrt{\lambda_i}\) (the “\(1\sigma\)” ellipse) and \(2\sqrt{\lambda_i}\) (the “\(2\sigma\)” ellipse), where \(\lambda_i\) are the eigenvalues of \(\Sigma\).
A subtlety worth highlighting. In one dimension the \(\pm 1\sigma\) and \(\pm 2\sigma\) intervals contain 68.3% and 95.4% of the probability mass. In two dimensions, the corresponding ellipses contain considerably less, because \(\Delta^2 \sim \chi^2_2\) and
\[
\prob(\Delta^2 \le k^2) = 1 - e^{-k^2/2}.
\]
Questions to consider:
What does “positive definite” mean and why is this a requirement for the covariance matrix \(\Sigma\)?
Answer
A symmetric matrix (such as a covariance matrix) is positive definite if all of its eigenvalues are greater than zero. This ensures that
the variance of any linear combination of random variables is non-negative;
that no variable is a linear combination of others (no collinear variables);
the covariance matrix is invertible.
What is plotted in each part of the graph (called a “corner plot”)?
What effect does changing \(\mu_1\) or \(\mu_2\) have?
What effect does changing \(\Sigma_{11}\) or \(\Sigma_{22}\) have? What if the scales for \(x_1\) and \(x_2\) were the same?
What happens if \(\rho_{12}\) is equal to \(0\) then \(+0.7\) then \(-0.7\).
What would happen if you were allowed to set \(|\rho_{12}| \leq 1\)? Explain what goes wrong.
So what characterizes independent (uncorrelated) variables versus positively correlated versus negatively correlated?
Follow-up on correlated posteriors
The first Gaussian 2D PDF in the last section is characterized by elliptical contours of equal probability density whose major axes are aligned with the \(x_1\) and \(x_2\) axes.
This is a signature of independent random variables.
The second Gaussian 2D PDF had major axes that were slanted with respect to the \(x_1\) and \(x_2\) axes.
This is a signature of correlated random variables.
Let’s look at a case with slanted posteriors and consider analytically (in the next section) what we should expect with correlations.
We’ll use the exercise notebook 📥 Parameter estimation example: fitting a straight line as a guide.
What are we trying to find? As in other notebooks, we seek \(p(\pars|D,I)\), now with \(\pars =[b,m]\) (you should review the statistical model).
Note
Some comments on the implementation:
Note that \(x_i\) is randomly distributed uniformly.
Log likelihood gives fluctuating results whose size depend on the # of data points \(N\) and the standard deviation of the noise \(dy\).
Note: log(posterior) = log(likelihood) + log(prior)
Explore!
Play with the notebook and explore how the results vary with \(N\) and \(dy\).
Observation about priors:
Observations about the second set of data:
True answers are intercept \(b = 25.0\), slope \(m=0.5\).
Flat prior gives \(b = -50 \pm 75\), \(m = 1.5 \pm 1\), so barely at the \(1\sigma\) level.
Symmetric prior gives \(b = 25 \pm 50\), \(m = 0.5 \pm 0.75\), so much better.
Distributions are wide (not surprising since only three points!).
Compare priors on the slope \(\Longrightarrow\) uniform in \(m\) vs. uniform in angle.
With the first set of data with \(N=20\) points, does the prior matter?
With the second set of data with \(N=3\) points, does the prior matter?
Recap: what does it mean that the ellipses are slanted? And why is it expected?
Answer
It means that the slope and intercept are correlated. Intuitively, if we want to maintain a good fit when we change the slope, we will need to change the intercept as well. They are not independent!
2D PDF with a quadratic approximation
Consider a two-dimensional log likelihood \(L(X,Y)\). We’ll analyze it in a quadratic approximation.
First, find the mode \(X_0\), \(Y_0\) (best estimate) by differentiating
\[\begin{split}\begin{align}
L(X,Y) &= \log p(X,Y|\{\text{data}\}, I) \\
\quad&\Longrightarrow\quad
\left.\frac{dL}{dX}\right|_{X_0,Y_0} = 0, \
\left.\frac{dL}{dY}\right|_{X_0,Y_0} = 0
\end{align}\end{split}\]
To check reliability, Taylor expand around \(L(X_0,Y_0)\):
\[\begin{split}\begin{align}
L &= L(X_0,Y_0) + \frac{1}{2}\Bigl[
\left.\frac{\partial^2L}{\partial X^2}\right|_{X_0,Y_0}(X-X_0)^2
+ \left.\frac{\partial^2L}{\partial Y^2}\right|_{X_0,Y_0}(Y-Y_0)^2 \\
& \qquad\qquad\qquad + 2 \left.\frac{\partial^2L}{\partial X\partial Y}\right|_{X_0,Y_0}(X-X_0)(Y-Y_0)
\Bigr] + \ldots \\
&\equiv L(X_0, Y_0) + \frac{1}{2}Q + \ldots
\end{align}\end{split}\]
It makes sense to do this in (symmetric) matrix notation:
\[\begin{split}
Q =
\begin{pmatrix} X-X_0 & Y-Y_0
\end{pmatrix}
\begin{pmatrix} A & C \\
C & B
\end{pmatrix}
\begin{pmatrix} X-X_0 \\
Y-Y_0
\end{pmatrix}
\end{split}\]
\[
\Longrightarrow
A = \left.\frac{\partial^2L}{\partial X^2}\right|_{X_0,Y_0},
\quad
B = \left.\frac{\partial^2L}{\partial Y^2}\right|_{X_0,Y_0},
\quad
C = \left.\frac{\partial^2L}{\partial X\partial Y}\right|_{X_0,Y_0}
\]
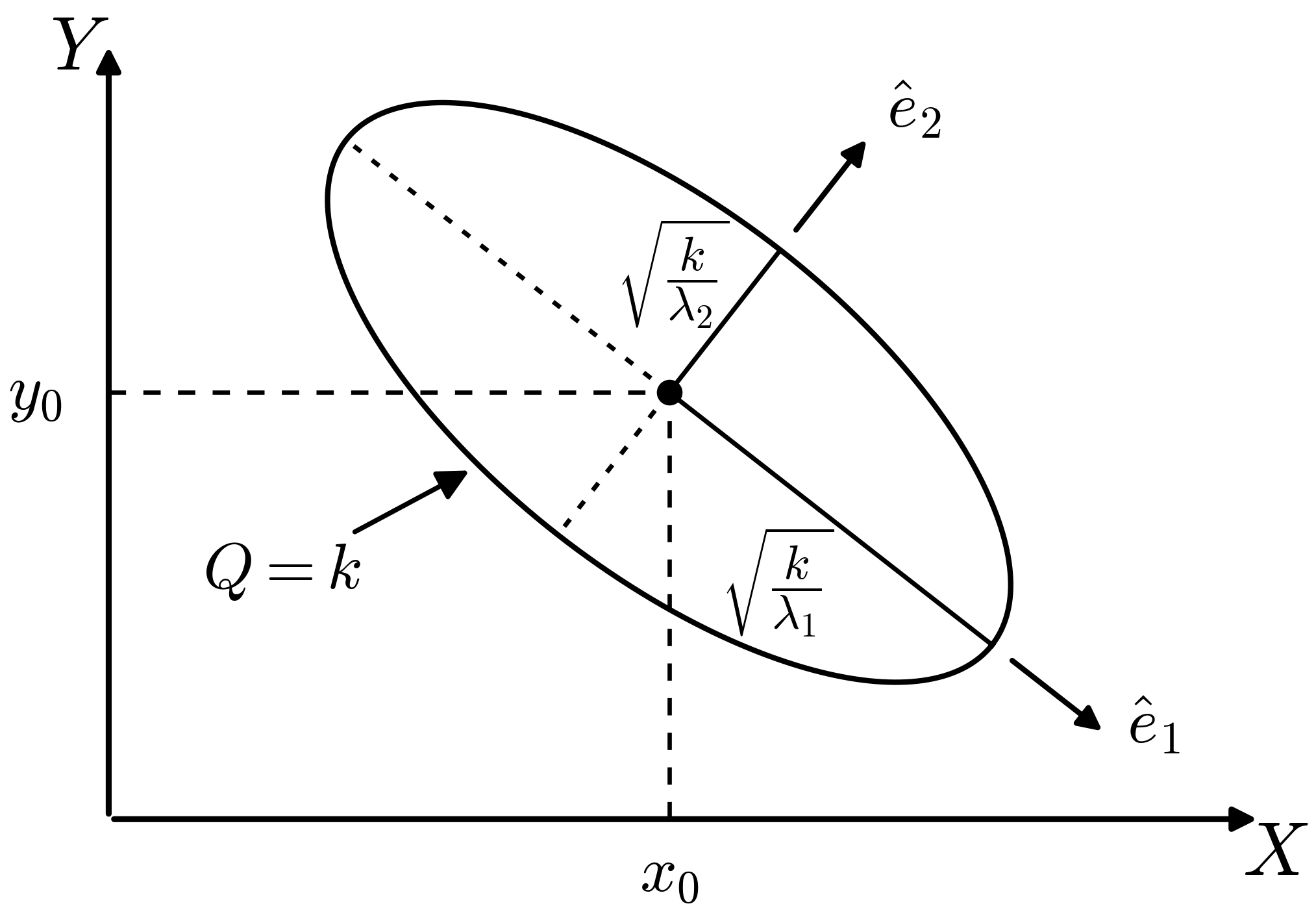
So in a quadratic approximation, the contour \(Q=k\) for some \(k\) is an ellipse centered at \(X_0, Y_0\) (as in the figure). The orientation and eccentricity are determined by \(A\), \(B\), and \(C\).
The principal axes are found from the eigenvectors of the Hessian matrix \(\begin{pmatrix} A & C \\ C & B \end{pmatrix}\):
\[\begin{split}
\begin{pmatrix}
A & C \\
C & B
\end{pmatrix}
\begin{pmatrix}
x \\ y
\end{pmatrix}
=
\lambda
\begin{pmatrix}
x \\ y
\end{pmatrix}
\quad\Longrightarrow\quad
\lambda_1,\lambda_2 < 0 \ \mbox{so $(x_0,y_0)$ is a maximum}
\end{split}\]
If the major and minor axes of the ellipse are aligned with the \(x\)-axis and \(y\)-axis (so \(C=0\)), the analysis is simple: the eigenvalues are \(A\) and \(B\) and the error-bars for \(X_0\) and \(Y_0\) will be inversely proportional to the modulus of their square roots.
What if the ellipse is skewed?
See Sivia section 3.3 [SS06] for a thorough treatment.
Compare Gaussian noise sampling to lighthouse analysis
Here we observe that Gaussian noise sampling is carried out just like the radioactive lighthouse analysis from exercise notebook 📥 Radioactive lighthouse problem (which we assume you have worked through).
Jump to the Bayesian approach in the exercise notebook 📥 Parameter estimation example: Gaussian noise and averages II.
The goal is to sample a posterior \(p(\pars|D,I)\)
\[
p(\mu,\sigma | D, I) \leftrightarrow p(x_0,y_0|X,I)
\]
where \(D\) on the left are the \(x\) points and \(D=X\) on the right are the \(\{x_k\}\) where scintillation flashes are detected.
What do we need? From Bayes’ theorem, we need
\[\begin{split}\begin{align}
\text{likelihood:}& \quad p(D|\mu,\sigma,I) \leftrightarrow p(D|x_0,y_0,I) \\
\text{prior:}& \quad p(\mu,\sigma|I) \leftrightarrow p(x_0,y_0|I)
\end{align}\end{split}\]
You are generalizing the functions for log PDFs and the plotting of posteriors that are in 📥 Radioactive lighthouse problem.
Note the functions for log-prior and log-likelihood in 📥 Parameter estimation example: Gaussian noise and averages II. Here \(\pars = [\mu,\sigma]\) is a vector of parameters; cf. \(\pars = [x_0,y_0]\).
Let’s step through the essential set up for emcee.
It is best to create an environment that will include emcee and corner.
Hint
Nothing in the emcee sampling part needs to change!
Basically we are doing 50 random walks in parallel to explore the posterior. Where the walkers end up will define our samples of \(\mu,\sigma\)
\(\Longrightarrow\) the histogram is an approximation to the (unnormalized) joint posterior.
Plotting is also the same, once you change labels and mu_true, sigma_true to x0_true, y0_true. (And skip the maxlike part.)
Maximum likelihood here is the frequentist estimate \(\longrightarrow\) this is an optimization problem. And you can read off marginalized estimates for \(\mu\) and \(\sigma\).
Question
Are \(\mu\) and \(\sigma\) correlated or uncorrelated?
Answer
They are uncorrelated because the contour ellipses in the joint posterior have their major and minor axes parallel to the \(\mu\) and \(\sigma\) axes. Note that the fact that they look like circles is just an accident of the ranges chosen for the axes; if you changed the \(\sigma\) axis range by a factor of two, the circle would become flattened.
Bottom line: the two analyses are completely analogous.